pFind Studio: a computational solution for mass spectrometry-based proteomics
Introduction
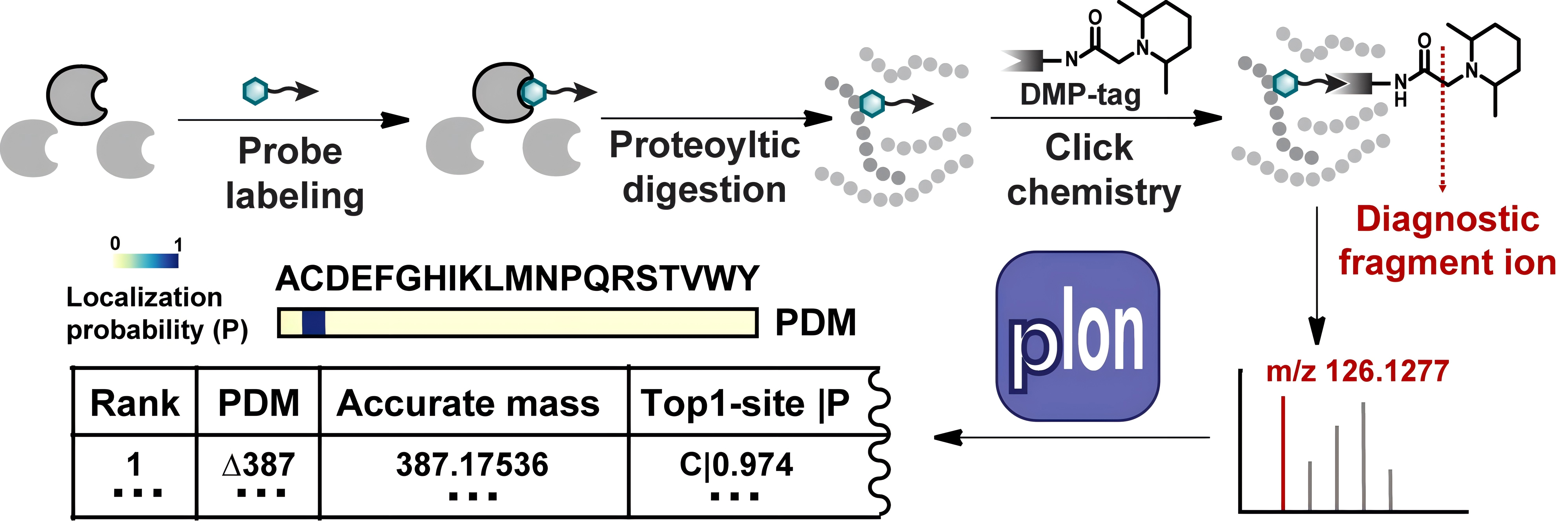
The ever-growing field of covalent drug discovery and chemoproteomics has fueled a need for bioconjugation methods with high selectivity in a native context. Given that the sheer number of functional groups present, achieving residue-specificity in biological systems remains a challenge. In order to identify the best residue-specific bioconjugation method, numerous small molecule models and/or purified proteins have been employed for rigorous method validation in terms of selectivity, efficiency and stability. Such in vitro experiments therefore became increasingly time consuming, yet they were unable to enumerate all functional groups present in complex, native biological systems. pIon is a computational tool that enables a cost-efficient pipeline for high-throughput evaluation of residue-specific bioconjugation chemistries. It starts with a rapid experimental phase, in which the proteome conjugated by a reactive probe is chemically coded for generating diagnostic report ions in tandem mass spectrometry analysis. The resulting MS data can be directly imported into pIon, which automatically calculates the accurate modification masses derived from a tested probe as well as the corresponding residue preferences. Thus, pIon has the potential to become a valuable option for routine evaluation of bioconjugation chemistries, thereby driving the field of bioconjugation chemistry to unprecedented dimensions and interfacing the worlds of biological and synthetic chemistry.
Downloads
Notice: Nov. 27, 2024 - pIon is currently available for free use. Click to download.
For demo data, please refer to Click to download..
For source code, please refer to github.
For detailed usage, please refer to user guide.
For other issues, please contact pengyaping21s@ict.ac.cn.